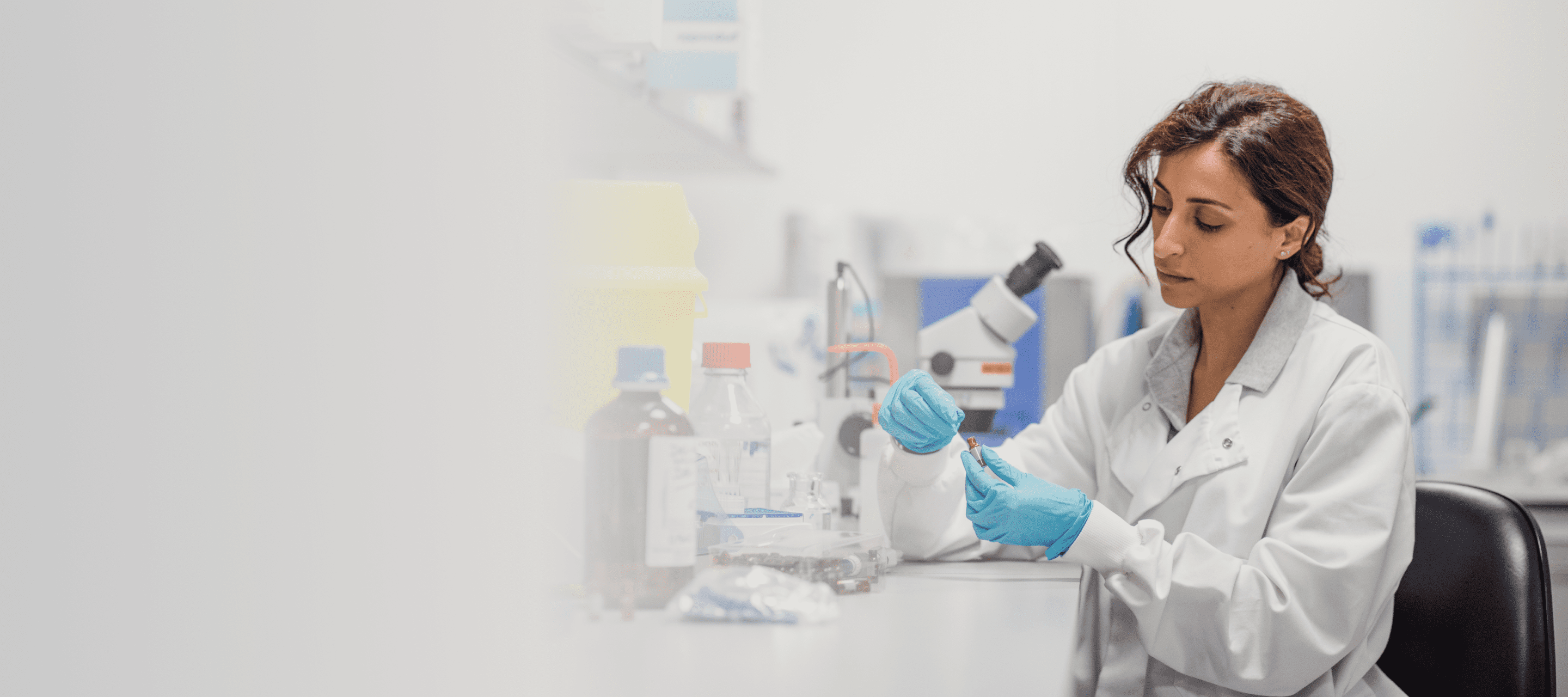


Improving efficiencies in UK diagnostics: An expert’s view
We talk to David Wells, CEO of the Institute of Biomedical Scientists, about how to improve efficiencies across the UK diagnostics sector
In many ways, the UK’s diagnostics sector is world-leading. For instance, the advancements that the country has made in genomics over the last few years has transformed the industry, as demonstrated by the role the UK played in sequencing COVID-19.
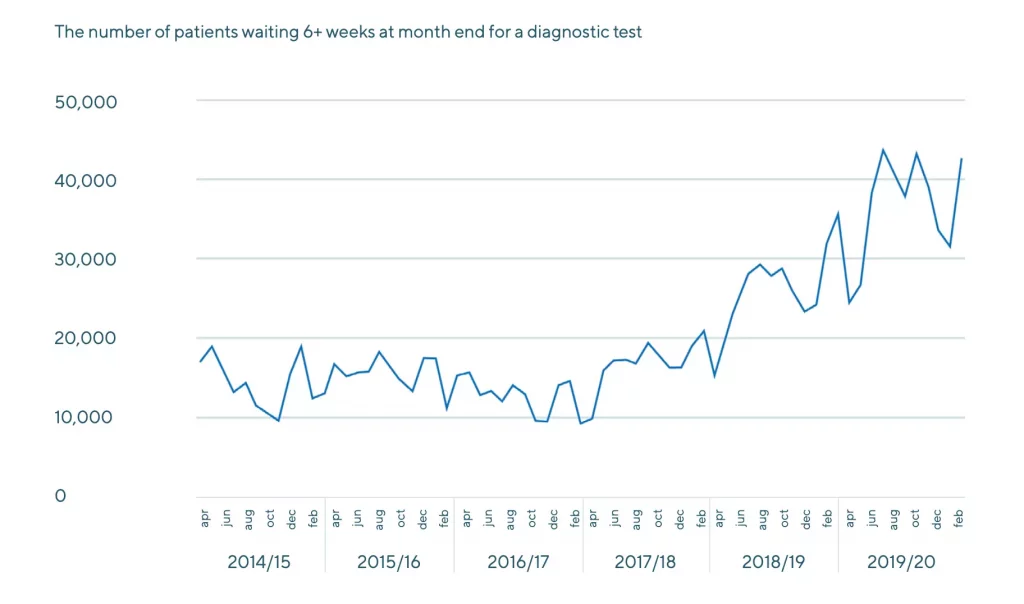
Yet, at the same time, the sector is plagued by inefficiencies. Long diagnostic waiting lists – exacerbated by COVID-19 – have created a pandemic of their own. Patients today are presenting with more advanced conditions as a result of these delays.
Increasing efficiencies within diagnostics is critical to preparing the healthcare sector for the challenges that lie ahead. A more efficient and proactive service is also key to saving money, resources and lives over the coming decade.
We sat down with David Wells, CEO of the Institute of Biomedical Scientists (the body that covers the training, education and support of Biomedical and Clinical Scientists), to talk about improving efficiency within the UK’s diagnostics sector. He spoke to us about the challenges today, the impact genomics will have, and how technology can help.

From ‘reactive’ to ‘proactive’: Diagnostics in the UK today
“The way that we run diagnostics in the UK has always been quite reactive,” Wells explains. “Although we’ve got world-leading screening programmes, medical facilities and funding for our NHS, we don’t always use that money wisely. And that means we have one of the worst cancer outcomes in the world. It also means we have long waiting lists for patients and, as a result, conditions that could have been treated with routine or simple interventions end up needing far more complex interventions.”
Wells gives the example of ovarian cancer, the sixth most common cancer in British women.
“In the UK, we have a really poor outcome for ovarian cancer patients. This is because the symptoms – bloating, pain, memory loss – are quite nonspecific, so are overlooked for months or years at a time. Women won’t get diagnosed until they present in A&E with an obstructive abdominal mass – at which point they have stage three or four ovarian cancer, which has a very, very poor outcome.
“If we had a more proactive diagnostic system in the UK, we could order tests earlier, or more regularly, and that cancer would be caught earlier. Sure, it would cost more each year, but it would save on tens of thousands of pounds spent treating a patient when they are at a later stage. And many more lives would be saved.”
He adds: “If we are able to increase efficiencies in diagnostics, we will be able to deliver a much cheaper and more effective service that also saves lives.”
“This comes down to the fact that the UK believes in personal responsibility for health,” explains Wells. “However, in Japan, there is more of an attitude of community responsibility. For example, companies can be fined for having overweight employees, or for not having a health initiative in place. If there is a particular community of people who are suffering from a particular condition, they are heavily monitored for those conditions.
“As a result of this emphasis on maintaining better health and screening more regularly, health is better in Japan, even though the country spends less on it.
“Of course, the caveat is that there is a difference in genetics between the two countries. But that can’t explain a 10-year difference in life expectancy,” he concludes.
How will the genomics revolution supercharge patient outcomes?
Genomic sequencing will play a significant role in the way diagnostic services are delivered in the UK over the coming years. As a result, the NHS has committed to sequencing 1 million genomes – 500,000 genomes in the NHS and 500,000 in UK Biobank – in order to improve health outcomes.
We ask Wells how he anticipates genomic sequencing will change diagnostic efficiency.
“Genomics will create massive efficiencies across the board in diagnostics,” he explains. “In the UK, we lead the world in genomics. Just look at how we have sequenced half the entire global genomics for the COVID-19 virus.
“Genomic sequencing gives us insight into the genetic causes of disease, so we can identify the people who are born with genetic predispositions for a certain medical condition,” he goes on. “Using genomics enables us to be proactive with patients who have a higher likelihood of developing a disease. So say we had a patient with the BRCA gene, we know we have to monitor them more often for ovarian cancer or breast cancer. As a result of this intervention, they will be more likely to live for longer, and remain a taxpayer who contributes to the NHS.”
Genomic sequencing also allows for more targeted, personalised treatment options.
“A better understanding of genetics helps us understand why some cancers don’t respond to certain treatments,” says Wells. “Let’s look at lung cancer, for example. When we started looking at lung cancer as a diagnostic step, around 20-30 years ago, we used to look through a light microscope and were able to identify four different types of lung cancer. Your treatment would depend on which of those four lung cancers you were deemed to have.
“Once we started looking at the proteins in those cancer cells, we realised there were more types, and identified around 26 genetic causes. Using this approach, we could target more specifically than before.
“When we started looking at the genomics of lung cancer, however, we found that there were far, far more types of cancer than previously thought. For just a third of those cases of cancer, we found that there are thousands of genetic causes,” he explains. “We realised that those genetic mutations also occur in the bowel, brain, bones, etc. And that if you have the genetic mutation for bowel cancer, even if it’s presenting as lung cancer, you can use the same treatment you would for bowel cancer and it works.”
Proteomics, and the future of diagnostics
“The truly exciting thing about genomics is how it will act as a stepping stone towards the real game changer: Proteomics,” says Wells.
“The problem we’ve got at the moment is that not all our genes switched ‘on’. We all have genes from our mothers and genes from our fathers, and some of them are switched ‘on’, while some are switched ‘off’. Using genomics to discover a mutation in a person’s tumour means making the assumption that that gene is switched on. But if it’s not, the targeted treatment you give them may not help.
“Whereas proteomics looks for the genes which are active or not active. It is much cheaper because it uses mass spectrometry, and you can actually start looking for the proteins that are expressed by that mutation,” he explains. “This could be established with a very simple blood test that the lab can then scan for the proteins that are suggestive of a mutation, and investigate where that cancer is. If found, you can start a treatment plan that responds to that specific mutation.
“This could shorten the cancer pathway, turning it from a drawn-out process where it takes two weeks to even see a doctor, to a system where you have a treatment plan before the condition even presents itself.“
What are the barriers for genomics today?
Wells argues that what is holding the UK’s diagnostics sector back today is mostly capacity issues.
“What we have is a brain and bandwidth problem. We don’t have enough people able to interrogate and understand what the genome is telling us,” he says.
“So with those thousands of different genomic mutations for lung cancer, we don’t have thousands of people to look at each mutation, which would be ideal. We have a much smaller number, looking at a whole gambit of genetic mutations.
“There is limited time for thinking, research and development at the moment. We need more highly-trained scientists who work in this environment to help pin down all the genomic mutations that are present,” Wells says.
“We also need to adopt technology which will help us combat what we lack in manpower. The ability to use machines to run research labs 24/7 would massively increase the speed and scale at which genomic sequencing could take place. Being able to leave a machine running overnight would mean doubling the capacity of a lab.”
That is why he argues that automation is a critical component of ushering in the genomics revolution.
Why automation holds the key to unlocking genomics
With automation enabling labs to run 24/7, they will be able to conduct genomic sequencing at the scale required for the UK to reach its goal of sequencing out 1 million genomes. The technology also ensures data collection is far more accurate. These factors combined will usher in what Wells calls a ‘diagnostic data revolution’.
“There is no way a human today can interpret all of the data that comes out in the diagnostic fields,” Wells explains. “But with automation technologies, labs can harness the computational power to interpret the data.”
Automated equipment is key for labs to keep up with the scale of diagnostics today, he argues. Many labs start with partial automation, such as a single liquid handler or plate reader, and move towards integrating laboratory workflow automation, where multiple pieces of equipment are linked using robotics. The more automated a workflow is, the more efficient, accurate and fast a lab can run, and automation partners such as Automata are able to integrate laboratory workflow automation with existing lab instruments, or design entirely new ones to solve whatever problems a lab may have.
“What automation allows us to do is use our current capacity efficiently, and our current workforce efficiently,” Wells concludes. “That really is why it’s a game changer.”
Learn more about Automata’s automated genomic sequencing solution

From diagnosis to treatment, new technologies are revolutionising scientists’ approach to cancer…
Read more Automation in cancer genome sequencing
Automata and Illumina discuss the evolving landscape of genomics and its transformative…
Read more In conversation with Illumina: genetics in healthcare
Watch Francis Crick Institute’s Jérôme Nicod discusses the impact of automating genomic…
Read more On-demand webinar: Addressing liquid handling bottlenecks with automation